If you want to try out linux (Ubuntu), this is BY FAR the easiest method. Just download this installer. The next time you reboot you can choose Windows or Linux:
http://wubi-installer.org/index.php
Basically, a file (6GB by default) is allocated inside of windows and a boot loader is installed. On reboot, you can choose windows or linux. When you are done, you can uninstall it and it goes away. Or, there are ways to migrate that file into a fresh partition if you want.
Very smooth.
Thursday, January 31, 2008
Monday, January 28, 2008
R help in a browser
Really annoying to try to look up docs in a window you are using. Instead, try this:
help.start() # this will start up documentation in a browser
help(plot) # shows documentation in browswer window now!
data.frame in R (the key object)
I've been wading through R trying to figure out how it all works. It looks like the most important data structure is the data frame, and here I demo how to make a dataframe that a person can then play with. Yes, you can import data from a spreadsheet, but a person should understand the underlying data structure.
A data.frame is just a list of vectors or matrices where each has the same number of rows (think of the vectors as columns). It really is just the same as data in a spreadsheet:
In fact, when one reads in this type of info, one gets a data.frame back:
So, assuming we have some data like the above, now lets make it in R:

A data.frame is just a list of vectors or matrices where each has the same number of rows (think of the vectors as columns). It really is just the same as data in a spreadsheet:
| var1 | var2 | var3 | fac1 | fac2 |
|---|---|---|---|---|
| 1 | 3 | 4 | hot | TX |
| 0 | 2 | 3 | cold | WI |
| 3 | 8 | 2 | cold | UT |
In fact, when one reads in this type of info, one gets a data.frame back:
df = read.csv(".csv", quote="") # <-reads an unquoted csv file with headers
So, assuming we have some data like the above, now lets make it in R:
> var1 <- c(1,0,3) # '<-' is the assignment operator (like a directional '=')
# 'c()' is the concatenate operator (makes lists of things)
> var2 <- c(3,2,8)
> var3 <- array( c(4,3,2), c(3) ) # here's how you make a more mathematical vector
> fac1 <- c("hot", "cold", "cold")
> fac2 <- c("TX","WI","UT")
# alright, lets add all this stuff together into a dataframe:
> df <- data.frame(var1,var2,var3,fac1,fac2, row.names=NULL, check.rows=TRUE)
# creates this:
var1 var2 var3 fac1 fac2
1 1 3 4 hot TX
2 0 2 3 cold WI
3 3 8 2 cold UT
# want to see the names of the columns in the dataset:
> names(df)
[1] "var1" "var2" "var3" "fac1" "fac2"
# change the names:
> names(df) <- c("height", "weight", "nose.length", "temp", "state")
> df
height weight nose.length temp state
1 1 3 4 hot TX
2 0 2 3 cold WI
3 3 8 2 cold UT
# so, now we have a data.frame, the preferred data structure in R. Let the magic begin:
> plot(df)

Friday, January 25, 2008
Installing and basic plotting in R
Windows users:
http://cran.r-project.org/bin/windows/base/
Download R-2.6.1-win32.exe and install it like any program.
Start it like any other program from the start menu
Linux users:
sudo apt-get install r-base
Just type R in the terminal
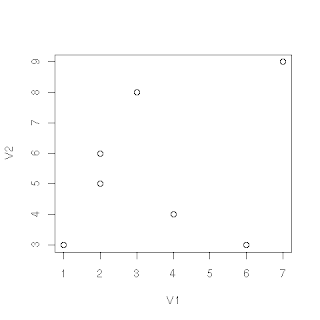
Read in a file
Go into excel or even a text editor and create a simple data set like this:
1,3
2,5
2,6
3,8
4,4
6,3
7,9
Where the second column is after the comma.
Save your file as a .csv
Look up where you saved it and insert that as the location in the following code:
data<-read.table("/home/eldon/Desktop/data.csv",sep=",") #this reads in your data and assigns it to the data frame "data"
plot(data) #this gives you a simple plot of your data
If you want to learn more about simple plotting just enter:
help(plot)
Thursday, January 24, 2008
Scientists for Better PCR
http://bio-rad.cnpg.com/lsca/videos/ScientistsForBetterPCR/
Good for a laugh
Good for a laugh
Wednesday, January 23, 2008
R from ruby
rsruby lets you run supposedly any function in R from ruby.
Elaborating on the sci-ruby page
I thought this was necessary, but don't so much any more:
Elaborating on the sci-ruby page
sudo apt-get install r-base ruby1.9.1-full ruby1.9.1-dev
sudo ln -s /usr/lib/R/lib/libR.so /usr/lib/libR.so
sudo ln -s /usr/share/R/include/R.h /usr/include/R.h
gem install rsruby -- --with-R-dir=/usr/share/R
I thought this was necessary, but don't so much any more:
# add a line to your .bashrc file to point to R:
echo "export R_HOME=/usr/lib/R" >> ~/.bashrc
source ~/.bashrc # read the bashrc file to get the variable
sudo make me a sandwich

http://imgs.xkcd.com/comics/sandwich.png
If you don't know what this is about, stick around and you might. eventually. after an explanation.
If you don't know what this is about, stick around and you might. eventually. after an explanation.
Subscribe to:
Posts (Atom)